Once you’ve sample your field, had it tested, and you have soybean cyst nematode (SCN), you’ll want to consider SCN-resistant varieties, but there are certain populations of SCN that can reproduce on certain SCN-resistant varieties so how do you know what resistant variety is best for your field – you find out the HG Type of your population – a costly test that UT is offering for free this year to TN farmers.
Previous articles at news.utcrops.com have discussed ‘What’s the difference between nematodes’, the silent yield robbers (‘Soilborne Pathogen Screening…’), and the ‘Free Soil Testing for Pathogens’ – how to sample, where to send sample, and the pathogens that are being screened. This article discusses the importance of SCN and the ‘HG Type’ test, where the HG stand for Heterodera glycines (the scientific name of soybean cyst nematode) and why it is the updated name of the previous ‘Race’ test. The majority of the information presented in this article came from TheSCNcoalition.com/resources (The Biology and Management of SCN).
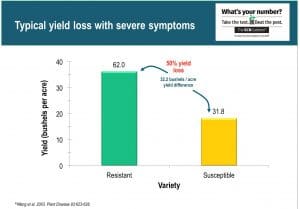
Why SCN is important – it can have significant impact on yield. Typical yield loss with severe symptoms (yellowing leaves and stunting) can be 50% or more based on the difference between growing a SCN susceptible variety and SCN resistant variety (Figure 1). Even with no symptoms, one can expect typical yield loss of 10%. So when it comes to planting soybean in an SCN infested field it should be a no brainer decision to plant a resistant variety, looking at the photo in Figure 2, which illustrates how “black and white or in the photo, dark green vs pale green… when comparing a susceptible variety on the left to resistant variety on the right. The bargain of the century here is that the seed of a resistant variety doesn’t cost any more than the seed of a susceptible variety – it’s free.” And on top of getting the yield benefit from a resistant variety, there’s a second payment in the nematode control that it will provide.

Now, how should one go about choosing which SCN resistant variety they should use? This is where the HG Type test comes into play, but first one needs to understand the biology of SCN resistance:
- Soybeans are resistant by containing several resistance genes (called Rhg genes, which stands for Resistance to Heterodera glycines – the scientific name for SCN)
- There are 7 main sources of known resistance the breeders use to create a SCN resistant variety (Indicator lines, listed in order: PI 548402 (Peking), PI 88788, PI 90763, PI 437654, PI 209332, PI 89772, and PI 548318 (Cloud)); but there are hundreds of others that plant geneticists and breeders have identified
- Furthermore, it has been found that it’s not just the presence of resistance genes but also the number of copies of the set of genes that determine how resistant the variety can be
- The scientific definition of a resistant soybean variety (there is no legal definition in the US) is that a resistant variety should allow less than 10% reproduction relative to a susceptible variety (in other words, there should be a 90% suppression or control)
The HG Type Test – provides information on the virulence of the SCN population and what resistant sources will work best
So the tool we have to test if a SCN population can reproduce more than 10% (relative to a susceptible variety) on any of the 7 main sources of resistance is called the HG Type test. The HG Type test replaces the SCN race test, which was devised in the late 1960s and used widely until the late 1990s. The reason for the switch is that the race concept was shown to be inadequate and inaccurate to describe and predict SCN reproduction on resistant varieties. To conduct the HG Type test, the SCN population from a field is usually increased in a greenhouse on a susceptible variety for 3 to 4 weeks. Then the SCN population is divided up across “HG type indicator” soybean lines (the 7 main sources of known resistance, mentioned previously, plus a susceptible check). The results are much more straight forward and easy to interpret compared to the old race test. For example, a HG type 2.4 SCN population has elevated reproduction on indicator lines 2 and 4 (2 = PI 88788 and 4 = PI 437654). SCN populations that do not have ≥10% reproduction on any HG type indicator line are HG type 0.
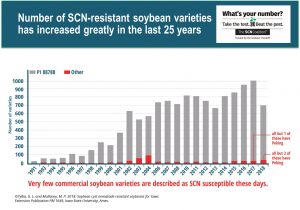
Now over the years there has been more and more SCN resistant commercial varieties released (~1,000 varieties in 2017 compared to only ~100 in 1997), but looking at the source of resistance in the majority of those varieties is alarming (Figure 3). The grey color in each bar, in Figure 3, are varieties with the PI 88788 source of resistance, showing that very few varieties have any of the other 6 main sources of resistance or the hundred or more others that are known. The main reason for this trend goes back to “yield ability” – there has been a “yield drag” or lowering the yield potential that had resulted from introducing these other SCN sources of resistance into a soybean variety. It has been difficult to get both good SCN resistance and high agronomics in a soybean variety using any source of resistance other than PI 88788. Hence, ~97% of varieties farmers can pick from have the PI 88788 SCN resistance. Remember these varieties can still differ from each other due to gene copy number, but it still spells certain doom for the longevity of the PI 88788 resistance (simply put in this entertaining ~2 min YouTube video: How the soybean cyst nematode (SCN) problem evolved).
Can you image what would happen to weed populations if a single herbicide active ingredient was used over and over for 25+ years? The exact same thing happens with extended use of SCN resistance genes. Research from Iowa, has shown populations of SCN are starting to reproduce at levels of 50-60% or more on PI 88788 (Figure 3). Hence, more varieties are needed with a diversity of SCN resistance that can be rotated in fields with SCN to maintain efficacy of the genetic resistance in soybean and avoid SCN populations developing resistance.
In Tennessee, we don’t know if we are in the same situation as Iowa or if we are still at a stage to try to avoid the “train wreck in slow motion” that is being observed in other soybean growing areas in regards to SCN populations reproducing on the PI 88788 resistance. So what can you do to help – soil sample your field, have it tested – FOR FREE – information on sampling and where to send can be found at UTcrops.com –> Soybean –> Diseases& Nematodes –> at bottom of publication section, Nematode Sampling Guide and Submission Form. If SCN is found in your sample, UT is also offering free HG Type testing (when able from sample) so we can better understand what is going on in our soils and go forward with the best management practices. Funding for the free processing of samples is provided by the TN Soybean Promotion board and the United Soybean Board. More information about SCN, as well as individual state resources can be found at TheSCNcoalition.com.

